Tissue dissociation is the fundamental step that enables researchers to isolate viable cells from solid tissues for downstream biological analysis.
Modern life-science workflows — including single-cell sequencing, flow cytometry, organoid culture, and drug discovery — all depend on the ability to convert complex tissues into high-quality single-cell suspensions.
However, isolating cells from tissue is not trivial. Different tissues require different dissociation strategies, and the chosen method can strongly influence cell viability, surface marker preservation, and experimental reproducibility.
This guide provides an overview of tissue dissociation methods, compares enzymatic and mechanical approaches, and explains how researchers can select the most suitable strategy for their specific applications.
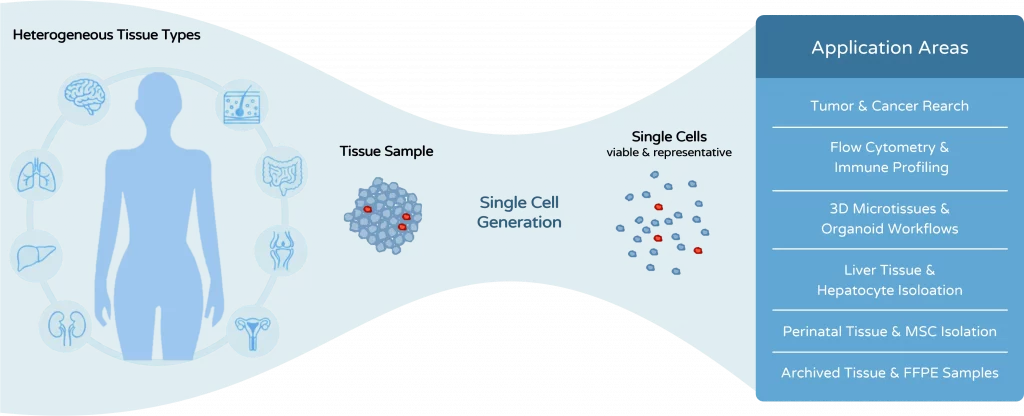
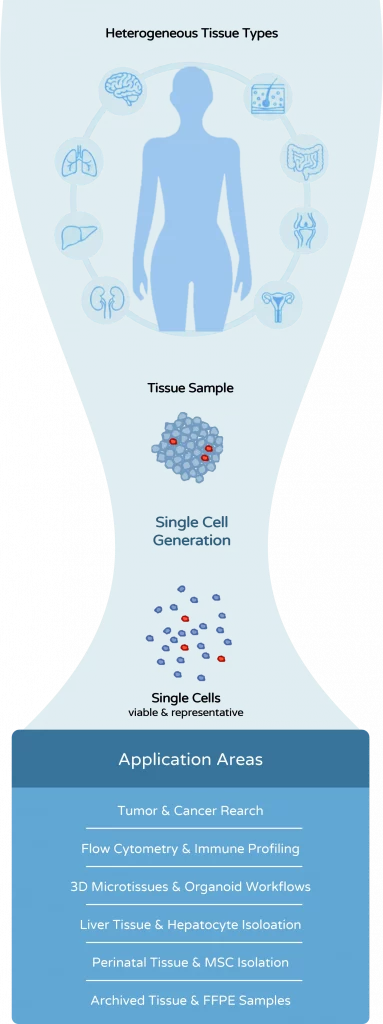
The dissociation method is not merely a preparatory step. It directly determines cell quality, experimental reliability, and the success of downstream analyses.
Tissue dissociation is often treated as a preparatory step in biological workflows. In reality, it is one of the most critical determinants of experimental quality. The way tissue is processed directly affects cell viability, cell phenotype, and the reliability of downstream analyses
Tissue Type:
The inherent characteristics of a tissue – including density, extracellular matrix composition, and cellular fragility – strongly influence the optimal dissociation approach. Delicate tissues such as brain often require gentler methods, whereas dense tissues such as cartilage or fibrotic tumors may require stronger mechanical disruption or enzymatic digestion.
Extracellular Matrix Density:
Tissues rich in collagen, elastin, or other ECM components often require enzymatic digestion targeting these structures. In tissues with looser extracellular matrices, mechanical dissociation alone may be sufficient to release cells.
Downstream Analysis Requirements:
The intended application determines the required cell quality. Techniques such as flow cytometry or single-cell sequencing require preservation of sensitive surface markers and high cell viability, which may favor gentler or enzyme-free dissociation strategies.
Regulatory Context:
Clinical or translational applications – for example in cell therapy manufacturing – often require highly reproducible, standardized, and GMP-compatible workflows to ensure safety and consistency.
Reproducibility and Throughput:
For high-throughput studies or multi-site collaborations, standardized and automated dissociation workflows can improve reproducibility and reduce operator-dependent variability.
By systematically evaluating these factors, researchers can select a dissociation strategy that aligns with both the biological properties of the tissue and the goals of the experiment.
Ultimately, the success of downstream analyses — including single-cell sequencing, flow cytometry, drug screening, and cell culture — depends strongly on the quality of the initial tissue dissociation step.
Recent advancements in tissue dissociation techniques for cell manufacturing, single-cell analysis, and downstream processing
Aaron Jankelow (2025) – Stem Cells Translational Medicine [↗︎]
Protocol for optimized mononuclear cell isolation from liver and tumor tissue using mechanical or enzymatic digestion
Sachin Chauhan (2026) – STAR Protocols [↗︎]
Long-term functional drug and immunotherapy screening in immune-competent patient-derived microtissues across brain tumors
Manon Bouzereau (2025) – Neuro-Oncology [↗︎]
Multiple strategies exist to isolate cells from tissues. The optimal dissociation method depends on tissue characteristics, extracellular matrix composition, and the requirements of the downstream experiment.
Suggested follow-up pages:
• /mechanical-tissue-dissociation/ [PLACEHOLDER]
• /enzyme-free-tissue-dissociation/ [PLACEHOLDER]
• /enzymatic-tissue-dissociation/ [PLACEHOLDER]
| Enzymatic | Manual Mechanical | Automated Mechanical (TissueGrinder) | |
|---|---|---|---|
| Principle | Chemical digestion of extracellular matrix proteins | Physical shear or cutting (mincing, trituration) | Programmable mechanical dissociation in a controlled system |
| Time | 30–120 min | 10–30 min | 2–6 min |
| Viability | Moderate–High | Low | Moderate–High |
| Reproducibility | Variable | Low | High |
| Enzyme Residue | Yes | None | None |
| Automation | No | No | Full |
| Impact on Cells | Risk of marker cleavage, phenotype changes, enzyme exposure | Operator-dependent shear stress, inconsistent recovery | Defined mechanical stress, potential impact on sensitive cells, partial loss of cellular integrity |
| Suitable For | Many tissues where surface markers are not critical | Small labs, exploratory workflows | Oncology, immune profiling, stem cells, GMP workflows |




Different dissociation strategies produce markedly different outputs, ranging from dispersed single cells to preserved multicellular fragments and microtissues.
Recent advancements in tissue dissociation techniques for cell manufacturing, single-cell analysis, and downstream processing
Aaron Jankelow (2025) – Stem Cells Translational Medicine [↗︎]
Protocol for optimized mononuclear cell isolation from liver and tumor tissue using mechanical or enzymatic digestion
Sachin Chauhan (2026) – STAR Protocols [↗︎]
Rebuilding the microenvironment of primary tumors in humans: a focus on stroma
Sung Ho Mun (2024) – Experimental & Molecular Medicine [↗︎]
Selecting the optimal dissociation strategy is critical for successful cell isolation and requires careful consideration of several biological and experimental factors. Researchers typically choose between enzymatic, mechanical, or hybrid approaches depending on the characteristics of the tissue and the requirements of the downstream workflow.
Tissue Type:
The inherent characteristics of a tissue – including density, extracellular matrix composition, and cellular fragility – strongly influence the optimal dissociation approach. Delicate tissues such as brain often require gentler methods, whereas dense tissues such as cartilage or fibrotic tumors may require stronger mechanical disruption or enzymatic digestion.
Extracellular Matrix Density:
Tissues rich in collagen, elastin, or other ECM components often require enzymatic digestion targeting these structures. In tissues with looser extracellular matrices, mechanical dissociation alone may be sufficient to release cells.
Downstream Analysis Requirements:
The intended application determines the required cell quality. Techniques such as flow cytometry or single-cell sequencing require preservation of sensitive surface markers and high cell viability, which may favor gentler or enzyme-free dissociation strategies.
Regulatory Context:
Clinical or translational applications – for example in cell therapy manufacturing – often require highly reproducible, standardized, and GMP-compatible workflows to ensure safety and consistency.
Reproducibility and Throughput:
For high-throughput studies or multi-site collaborations, standardized and automated dissociation workflows can improve reproducibility and reduce operator-dependent variability.
By systematically evaluating these factors, researchers can select a dissociation strategy that aligns with both the biological properties of the tissue and the goals of the experiment.
Enzymatic dissociation employs proteolytic enzymes to degrade extracellular matrix components and release cells from tissue structures. This approach is widely used when tissues contain dense connective matrices that physically restrict cell release.
Enzymatic digestion targets proteins, polysaccharides, and other extracellular components that bind cells together. Common enzymes used in tissue dissociation protocols include:
Collagenase:
Degrades collagen, the most abundant structural protein of the extracellular matrix. Different collagenase types (e.g., Type I or Type IV) are frequently used for tissues such as tumors, liver, adipose tissue, and pancreas.
Papain:
A cysteine protease often used for neural tissue dissociation because of its relatively gentle action, which can help preserve neuronal viability.
Trypsin:
A serine protease commonly used to detach adherent cells from culture surfaces. It can also be used for tissue dissociation but may cleave sensitive surface proteins if exposure is prolonged.
DNase:
Frequently added to digestion cocktails to degrade extracellular DNA released from damaged cells. This prevents cell clumping and improves the quality of single-cell suspensions.
high efficiency for connective or fibrous tissues
widely established protocols for many tissue types
effective for releasing cells embedded in dense extracellular matrices
enzyme batch variability may affect reproducibility
risk of surface protein degradation or marker cleavage
prolonged digestion can reduce cell viability
enzymatic exposure may alter cellular phenotypes or transcriptional profiles
requires washing steps that increase workflow complexity
Protocol for optimized mononuclear cell isolation from liver and tumor tissue using mechanical or enzymatic digestion
Sachin Chauhan (2026) – STAR Protocols [↗︎]
Suggested follow-up page:
• /enzymatic-tissue-dissociation/ [PLACEHOLDER]
Mechanical (or physical) dissociation separates cells from tissues using controlled physical forces rather than enzymatic digestion. These approaches disrupt tissue architecture through shear, cutting, or grinding forces, ensuring the biochemical integrity of the cell surface remains intact.
Mechanical dissociation encompasses a wide range of techniques designed to break tissue into smaller fragments and release cells while minimizing biochemical modification. Unlike enzymatic methods, which rely on chemical reactions, mechanical protocols use kinetic energy to overcome the cohesive forces of the extracellular matrix (ECM).
Manual Mincing or Chopping:
Using scalpels or surgical scissors, tissue is cut into sub-millimeter fragments. This increases the surface-to-volume ratio, facilitating either immediate cell release or preparing the sample for subsequent steps.
Shear-based Dissociation:
Tissue is forced through narrow orifices, specialized filters, or microfluidic channels. The resulting fluid shear stress gently pulls cells apart from the tissue matrix.
Grinding or Mashing:
Particularly effective for softer tissues (like spleen or lymph nodes), this involves using pestles, automated grinders, or mesh strainers to physically press cells out of the structural framework.
Pipetting or Vortexing:
Repeated aspiration (trituration) or high-frequency agitation can disperse small cell aggregates or partially dissociated fragments into a true single-cell suspension.
Enzyme-Free Processing:
Eliminates the risk of “off-target” digestion of sensitive surface markers and receptors.
Biochemical Integrity:
Minimal risk of chemical modification to cell components or Alterations in transcriptional profiles.
Speed:
Often significantly faster than enzymatic incubation, which can take 30–90 minutes.
Temperature Control:
Excellent compatibility with ice-cold processing to preserve temperature-sensitive analytes.
Reproducibility:
Manual methods are highly operator-dependent, leading to variability between labs.
ECM Constraints:
Highly fibrotic or dense tissues (e.g., cartilage) may show lower yields without chemical assistance.
Cell Stress:
Excessive or unoptimized mechanical force can lead to physical rupture of fragile cell types.
A step towards enzyme-free tissue dissociation
Stefan Scheuermann et al. (2019) – Current Directions in Biomedical Engineering [↗︎]
TissueGrinder, a novel technology for rapid generation of patient-derived single cell suspensions from solid tumors by mechanical tissue dissociation
Stefan Scheuermann et al. (2022) – Frontiers in Medicine [↗︎]
Investigation of the Effect of High Shear Stress on Mesenchymal Stem Cells Using a Rotational Rheometer in a Small-Angle Cone–Plate Configuration
Mario Mand (2024) – Bioengineering [↗︎]
Explore detailed mechanical dissociation workflows → PLACEHOLDER!
Hybrid dissociation protocols combine enzymatic digestion with mechanical processing to improve efficiency while limiting the drawbacks of either approach alone.
In many modern workflows, tissues are first mechanically minced to increase the surface area available for enzymatic digestion. A shorter or lower-concentration enzymatic step then softens extracellular matrix components, after which mechanical processing releases the cells into suspension.
increase overall cell yield
shorten digestion times
reduce enzyme exposure
improve the balance between efficiency and cell viability
Hybrid approaches are therefore commonly used for complex tissues that cannot be efficiently dissociated using only mechanical or only enzymatic methods.
Protocol for optimized mononuclear cell isolation from liver and tumor tissue using mechanical or enzymatic digestion
Sachin Chauhan (2026) – STAR Protocols [↗︎]
Enzyme-free tissue dissociation avoids proteolytic digestion and relies on controlled mechanical disruption to release cells while preserving sensitive surface markers, cellular viability, and native gene expression profiles.
Enzyme-free tissue dissociation represents an alternative strategy to traditional enzymatic digestion methods. Instead of using proteolytic enzymes to degrade extracellular matrix components, these approaches rely on controlled mechanical forces to release cells from tissue while avoiding chemical modification of cellular structures.
The main goal of enzyme-free cell isolation is to preserve the native cellular state as much as possible. By avoiding enzymatic activity, researchers can better maintain sensitive surface markers, gene expression profiles, and cellular viability.
Controlled rotor-stator interactions progressively fragment tissue without enzymatic digestion, helping preserve cell integrity and surface markers.
Avoiding enzymatic digestion can help prevent several common problems associated with enzyme-based workflows:
degradation of cell-surface receptors required for cell sorting or immunological studies
cell stress or apoptosis caused by prolonged enzyme exposure
alterations of transcriptional profiles that can obscure biological signals
variability introduced by enzyme batch differences or digestion conditions
Because of these advantages, enzyme-free dissociation approaches are particularly valuable when working with delicate cell types, rare cell populations, or experiments that require maximal preservation of cellular integrity.
In practice, modern enzyme-free dissociation methods typically rely on precisely controlled mechanical disruption to release cells while minimizing damage. These workflows can enable:
high-viability single-cell suspensions
preservation of stromal and immune cell compartments
recovery of ECM-containing microtissue fragments
sequencing-ready RNA material
primary cells suitable for culture or downstream functional assays
Suggested follow-up page:
• /enzyme-free-tissue-dissociation/ [PLACEHOLDER]
Recent advancements in tissue dissociation techniques for cell manufacturing, single-cell analysis, and downstream processing
Aaron Jankelow (2025) – Stem Cells Translational Medicine [↗︎]
A step towards enzyme-free tissue dissociation
Stefan Scheuermann (2019) – Current Directions in Biomedical Engineering [↗︎]
Rapid single-cell physical phenotyping of mechanically dissociated tissue biopsies
Despina Soteriou (2023) – Nature Biomedical Engineering [↗︎]
Single Cell Isolation from Surgically Resected Tissue Via Mechanical Dissociation Using TissueGrinder
Prama Pallavi (2024) – Single Cell Analysis. Methods in Molecular Biology [↗︎]
Tissue dissociation is used across many different research workflows – each with its own requirements for cell quality, viability, and reproducibility. Whether you are working with tumor samples, immune cells, liver tissue, or 3D models, the choice of dissociation strategy can significantly impact your results.
Explore the key application areas below to see how tissue dissociation is applied in different contexts and which approaches are most relevant for your workflow.
Viable single-cell suspensions and ECM-preserved microtissues from complex solid tumors for profiling, sequencing, functional assays, and screening workflows. Tumor dissociation is especially relevant where heterogeneous tissue architecture, stromal content, and fragile immune or tumor cell populations must be handled reproducibly.
Solid tumors contain heterogeneous cellular compartments including tumor cells, stromal cells, immune infiltrates, and dense extracellular matrix. Preparing representative samples from these tissues requires reliable tumor dissociation protocols capable of generating viable single-cell suspensions from solid tumors or biologically meaningful tumor fragments that remain useful for downstream screening or culture-based follow-up.
Traditional enzymatic digestion approaches can introduce variability, damage surface markers, or disrupt extracellular structures that are still relevant for downstream assays. Researchers therefore frequently explore mechanical or hybrid dissociation strategies when the aim is not only to release cells, but also to preserve tumor architecture, ECM-associated features, or sensitive subpopulations.
Tumor dissociation for single-cell RNA sequencing
Preparation of tumor samples for flow cytometry and immune profiling
Generation of tumor microtissues for drug screening and immunotherapy assays
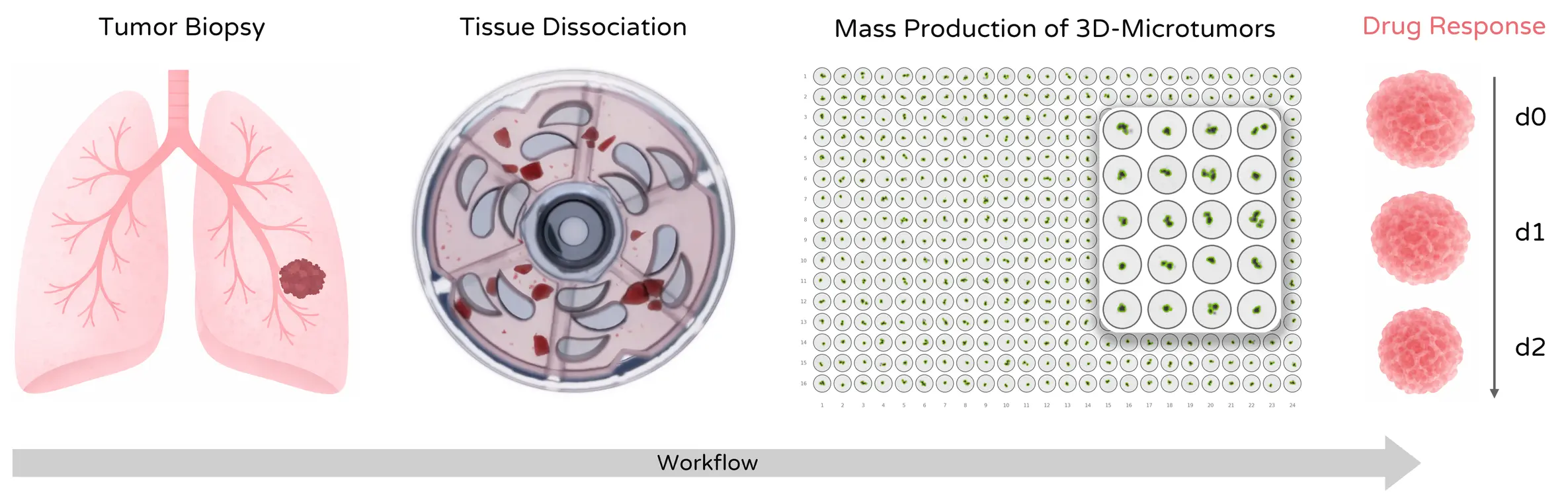
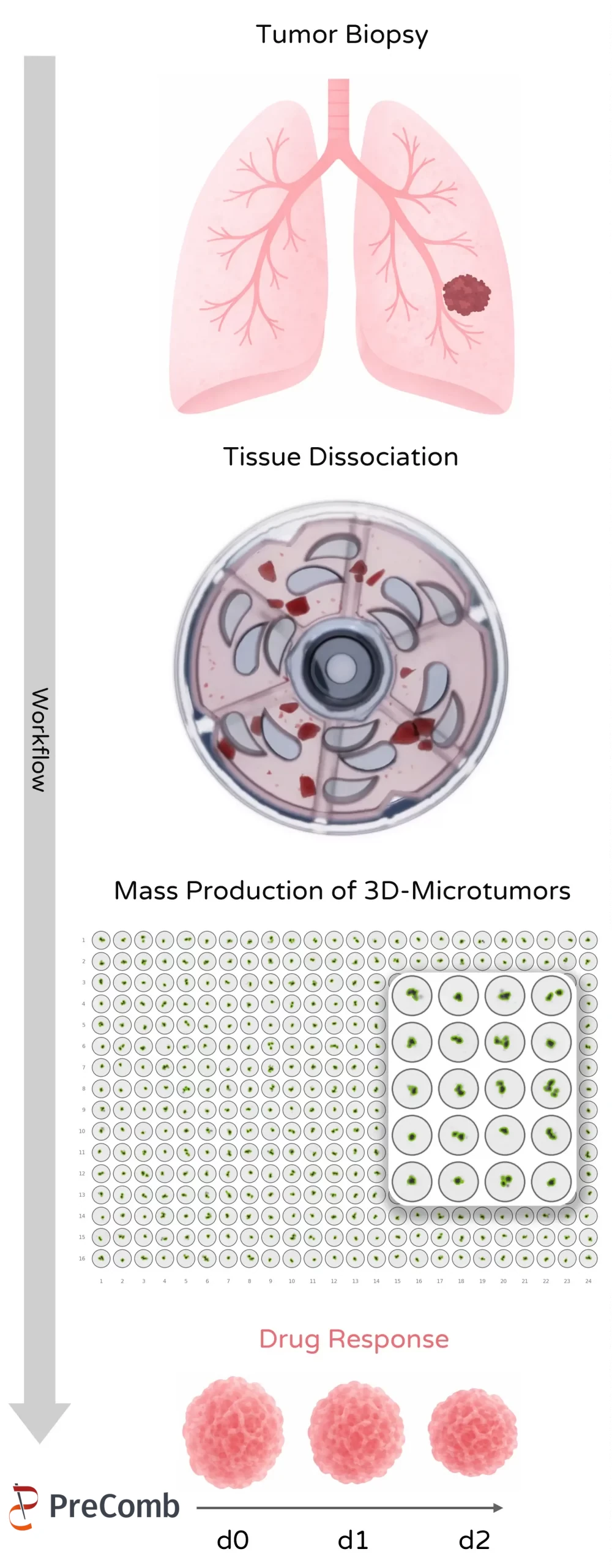
Mechanical tissue dissociation can preserve biologically relevant tumor fragments and support downstream microtissue generation for drug and immunotherapy testing.
Viable single-cell suspensions for molecular profiling and sequencing
ECM-preserved tumor fragments and microtissues for functional testing
Relevance across multiple tumor entities including HCC, glioblastoma, colorectal, pancreatic, lung, and metastatic tumors
Strong fit for the literature-backed areas already highlighted in the frozen-content concept
Tumor microenvironment analysis
Drug response testing
Tumor cell culture and cell line development
TissueGrinder, a novel technology for rapid generation of patient-derived single cell suspensions from solid tumors by mechanical tissue dissociation
Stefan Scheuermann (2022) – Frontiers in Medicine [↗︎]
Protocol for optimized mononuclear cell isolation from liver and tumor tissue using mechanical or enzymatic digestion
Sachin Chauhan (2026) – STAR Protocols [↗︎]
Long-term functional drug and immunotherapy screening in immune-competent patient-derived microtissues across brain tumors
Manon Bouzereau (2025) – Neuro-Oncology [↗︎]
Single Cell Isolation from Surgically Resected Tissue Via Mechanical Dissociation Using TissueGrinder
Prama Pallavi (2024) – Single Cell Analysis. Methods in Molecular Biology [↗︎]
Single-cell preparation from tissue for flow cytometry, FACS, immune profiling, and related analysis workflows. This application area is particularly important where preservation of surface epitopes, viability, and low debris are critical for reliable immunophenotyping.
Many laboratories perform tissue dissociation for flow cytometry in order to analyse immune or mixed cell populations directly from biopsies and fresh tissue samples. The quality of this preparation step strongly influences viability, debris levels, and preservation of the surface epitopes required for accurate immunophenotyping and downstream cell sorting.
Some enzymatic digestion protocols may cleave relevant markers or activate immune cells during processing. Researchers therefore look for workflows that support reliable FACS preparation from tissue while still producing clean, representative single-cell suspensions suitable for immune profiling, CyTOF, or subsequent sequencing workflows.


Automated mechanical dissociation can better preserve sensitive surface markers relevant for immunophenotyping than enzymatic tissue processing.
Immune cell isolation from tumor or inflamed tissue
Flow cytometry profiling of tissue-derived immune populations
Cell sorting prior to scRNA-seq or functional assays
Preservation of surface epitopes used in immunophenotyping
Reduced debris and clumping for more reliable acquisition
Compatibility with downstream single-cell and high-dimensional analysis
Strong relevance for marker-sensitive workflows such as CyTOF and immune profiling
Multiparameter flow cytometry
Immune profiling and immunophenotyping
Sorting for sequencing or proteomics workflows
Rapid single-cell physical phenotyping of mechanically dissociated tissue biopsies
Despina Soteriou (2023) – Nature Biomedical Engineering [↗︎]
Comparing the Influence on Surface Markers of Primary Isolated Splenocytes: Enzymatic Treatment vs. Mechanical Dissociation with TissueGrinder
E. Priebe (2023) – Fraunhofer scientific poster [↗︎]
Single cell secretome analyses of hepatic immune and stromal cells in liver disease
Richell Booijink (2024) – Dissertation at the University of Twente [↗︎]
Processing of fixed spheroids for TOBis barcoding, enzyme-free dissociation and antibody staining for CyTOF
Ralitsa R. Madsen (2024) – protocols.io [↗︎]
Preparation of viable tissue-derived material for 3D models, microtissues, organoid-related workflows, and drug screening applications. The focus here is on generating biologically useful cells or fragments rather than simply maximizing dissociation intensity.
Three-dimensional cell models such as spheroids, microtissues, and organoid-related systems are widely used in cancer research and drug discovery. Generating these models often begins with tissue dissociation steps that must deliver viable cells or fragments while preserving biological properties relevant for structured 3D culture.
Excessive enzymatic digestion can destroy extracellular matrix components and reduce the ability to form organized 3D models. Mechanical or carefully balanced hybrid workflows can therefore be valuable where researchers want to maintain ECM-associated features, generate tissue-derived microtissues, or support downstream organoid preparation and functional screening.
Compared with enzymatic digestion, mechanical dissociation can better preserve multicellular tumor fragments that subsequently form stable microtissues.
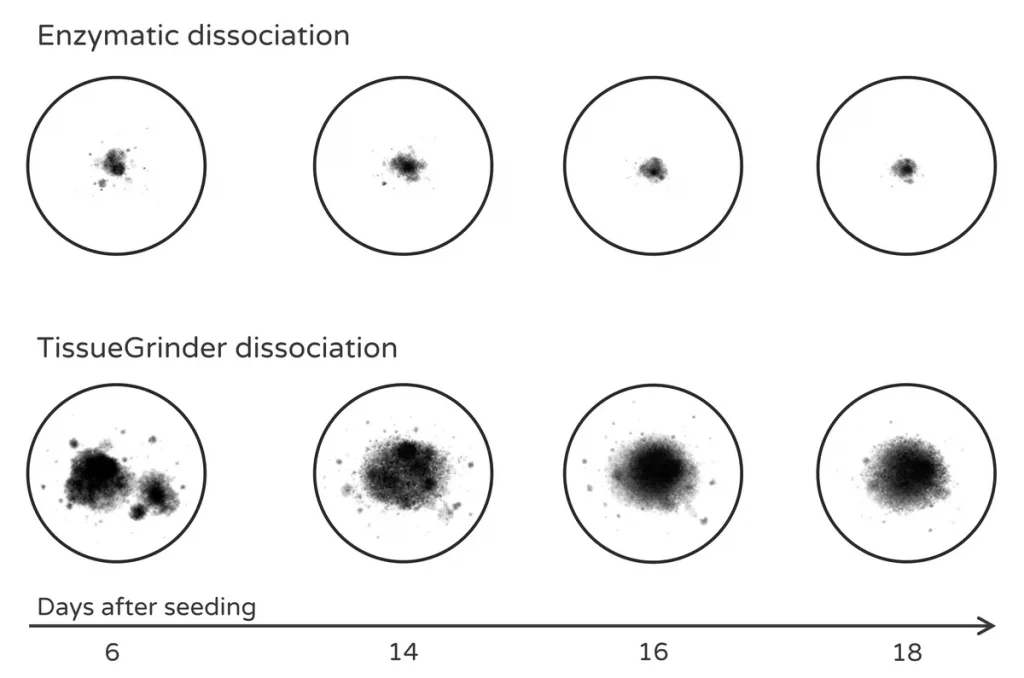
Preparation of tumor fragments for 3D microtissue drug screening
Generation of patient-derived organoid-related models
Formation of immune-competent tumor microtissues
Support for tissue-derived 3D model generation
Preservation of ECM features relevant for microarchitecture
Better fit for functional drug and immunotherapy assays than overly aggressive digestion
Drug response testing
Immunotherapy screening
Imaging-based phenotyping and multimodal analysis
Long-term functional drug and immunotherapy screening in immune-competent patient-derived microtissues across brain tumors
Manon Bouzereau (2025) – Neuro-Oncology [↗︎]
Oncogenic PIK3CA corrupts growth factor signaling specificity
Ralitsa R. Madsen (2024) – EMBO Molecular Medicine [↗︎]
SIGNAL-seq: Multimodal Single-cell Inter- and Intra-cellular Signalling Analysis
James Opzoomer (2024) – bioRxiv [↗︎]
Liver tissue dissociation for hepatocyte-related preparation, immune and stromal cell analysis, and extracellular vesicle workflows. Organ-specific liver applications broaden the hub beyond tumor-focused use cases and address an important set of analytical and cell-isolation questions.
Liver tissue contains hepatocytes together with immune, endothelial, and stromal cell populations embedded in extracellular matrix structures. Reliable liver tissue dissociation protocols are therefore required to prepare samples suitable for cellular and molecular analysis without unnecessarily compromising sensitive subpopulations or analytes of interest.
This application area is relevant not only for hepatocyte isolation but also for immune profiling from liver tissue, fibrosis and regeneration studies, and extracellular vesicle workflows that start directly from tissue-derived material. As a result, it helps the hub address important organ-specific search intents beyond oncology.
Liver dissociation for hepatocyte isolation
Preparation of immune cells from liver tissue
Extracellular vesicle studies derived from tissue material
Support for organ-specific dissociation workflows
Relevance for hepatocyte-related preparations and liver analysis
Connection to enzyme-free or reduced-enzyme analytical workflows
Liver disease research
Fibrosis and regeneration studies
EV characterization and molecular analysis
Comparative Assessment of Whole Organ Tissue Processing Methods for the Isolation of Extracellular Vesicles from Intact Organs
Mohammed Tayab Hussain (2025) – Journal of Extracellular Vesicles [↗︎]
Design, molecular characterization and therapeutic investigation of a novel CCR8 peptide antagonist that attenuates acute liver injury by inhibiting infiltration and activation of macrophages
Eline Geervliet (2025) – Acta Pharmaceutica Sinica B [↗︎]
Hepatocyte survival and proliferation by fibroblast growth factor 7 attenuates liver inflammation and fibrogenesis during acute liver injury via paracrine mechanisms
Eline Geervliet (2023) – Biomedicine & Pharmacotherapy [↗︎]
The development of hepatocellular carcinoma models in precision-cut tissue slices for therapeutic screening and precision medicine
Amy L. Collins (2024) – Dissertation at Newcastle University [↗︎]
Damaging mutations in LXRα uncouple lipogenesis from hepatotoxicity and implicate hepatic cholesterol sensing in human liver health
Sam M. Lockhart (2024) – bioRxiv [↗︎]
Umbilical cord and related perinatal tissues as sources for mesenchymal stromal cell isolation, culture expansion, and regenerative medicine research. This area is especially relevant where collagen-rich tissue matrices complicate standard processing.
Perinatal tissues such as umbilical cord represent an important source of mesenchymal stromal cells for research and translational workflows. Preparing these tissues typically involves specialized dissociation approaches capable of handling collagen-rich and elastic matrices such as Wharton’s jelly in a reproducible way.
This application area is important because researchers frequently search for umbilical cord dissociation and MSC isolation workflows that are more standardized than purely manual methods. It also adds a regenerative-medicine and stem-cell dimension to the hub, expanding the page beyond tumor and analytical use cases.
Umbilical cord dissociation for MSC isolation
Preparation of Wharton’s jelly fragments or suspensions
MSC culture and expansion workflows
Improved reproducibility in collagen-rich perinatal tissue handling
Strong relevance for MSC-focused culture workflows
Direct fit with Wharton’s jelly and umbilical cord search intents
Stem cell biology
Regenerative medicine research
Cell therapy development
Isolation of Porcine Umbilical Cord Cells by Mechanical Tissue Dissociation Using a Tissue Grinder
Katja Stange (2025) – Cells [↗︎]
Investigation of the Effect of High Shear Stress on Mesenchymal Stem Cells Using a Rotational Rheometer in a Small-Angle Cone–Plate Configuration
Mario Mand (2024) – Bioengineering [↗︎]
Hepatocyte survival and proliferation by fibroblast growth factor 7 attenuates liver inflammation and fibrogenesis during acute liver injury via paracrine mechanisms
Eline Geervliet (2023) – Biomedicine & Pharmacotherapy [↗︎]
Processing of archived and fixed tissue for RNA analysis, sequencing, and retrospective molecular workflows. This application area broadens the hub beyond viable-cell generation toward analytical sample preparation from clinically relevant archived material.
Archived clinical samples stored as FFPE tissue represent valuable resources for retrospective molecular analysis, but standard enzymatic digestion approaches are often ineffective for these materials. This makes FFPE tissue processing a distinct application area with different goals from fresh-tissue cell isolation.
Rather than focusing on viable cells, researchers in this area are often interested in preparing analyzable material for RNA extraction, sequencing, and other molecular pathology workflows. Including FFPE and archived tissue in the hub is therefore important for covering a broader range of clinically relevant search intents.

On the left, an example of single cells generated from FFPE samples with the TissueGrinder compared to a standard enzymatic dissociation on the right.
Above, an example of single cells generated from FFPE samples with the TissueGrinder compared to a standard enzymatic dissociation below.

Even archived FFPE samples can yield analyzable single-cell-like material when processed with enzyme-free mechanical dissociation.
Processing FFPE tissue for RNA or miRNA analysis
Preparation of archived tissue for sequencing
Dissociation of fixed tissue fragments for analytical workflows
Broader relevance for retrospective and archived-sample research
Connection to sequencing and molecular pathology use cases
Clear distinction from classic fresh-tissue dissociation workflows
RNA sequencing
Molecular pathology studies
Retrospective clinical analysis
Manual mechanical dissociation — such as mincing, grinding, or pipetting — can release cells efficiently, but the applied forces are difficult to standardize. Even small differences in handling can influence cell viability, yield, and phenotype preservation.
Automated tissue dissociation systems address these challenges by applying controlled mechanical forces under standardized conditions. Programmable dissociation profiles allow researchers to adapt mechanical processing to different tissue types while maintaining reproducibility between runs.
standardized mechanical parameters
improved experimental reproducibility
reduced operator-dependent variability
faster and more consistent processing
improved integration into regulated workflows
Automated mechanical dissociation platforms such as the TissueGrinder apply precisely controlled shear forces in a closed processing environment. This enables reproducible generation of viable cell suspensions while preserving sensitive cell structures and surface markers.
high-throughput research workflows
multi-center studies
clinical and translational applications
laboratories requiring standardized sample processing
Protocol for optimized mononuclear cell isolation from liver and tumor tissue using mechanical or enzymatic digestion
Sachin Chauhan (2026) – STAR Protocols [↗︎]
Single Cell Isolation from Surgically Resected Tissue Via Mechanical Dissociation Using TissueGrinder
Prama Pallavi (2024) – Single Cell Analysis [↗︎]
Tissue dissociation is the process of separating individual cells from solid tissue samples.
It involves breaking down the structural connections between cells – either mechanically, enzymatically, or through a combination of both – to generate a usable single-cell suspension.
This step is essential for many downstream applications, including flow cytometry, single-cell sequencing, and cell culture, where intact and viable cells are required for reliable analysis.
Mechanical and enzyme-free dissociation methods generally preserve cell-surface markers better than enzymatic approaches.
Proteolytic enzymes such as trypsin or collagenase can cleave or modify surface proteins, which may affect immunophenotyping or cell sorting results. In contrast, carefully controlled mechanical dissociation avoids chemical modification and helps maintain native marker expression.
The optimal method, however, depends on the tissue type and the specific markers of interest.
Common enzymes used for tissue dissociation include:
These enzymes are often used in combination to improve dissociation efficiency, especially in dense or fibrotic tissues.
Mechanical dissociation works well for many tissues, especially softer or moderately structured ones. However, it may be less efficient for highly fibrotic or dense tissues, such as skin or strongly collagen-rich tumors.
In such cases, hybrid approaches – combining mechanical processing with mild enzymatic digestion – can improve cell yield while still limiting enzyme exposure.
The choice of method should always be adapted to the tissue properties and the requirements of the downstream application.
Cell viability during tissue dissociation can be improved by optimizing several key factors:
Carefully balancing efficiency and gentleness is critical. Overly harsh conditions – whether mechanical or enzymatic – can reduce viability and affect downstream results.

Experience reproducible cell isolation with the TissueGrinder.
Transform complex tissue into high-quality single cells in minutes while preserving cell viability and surface markers. Standardize your workflows and reduce variability in tissue processing.